This is the Linux app named Linking Yabi with RDA whose latest release can be downloaded as javadoc.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named Linking Yabi with RDA with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
Linking Yabi with RDA
DESCRIPTION
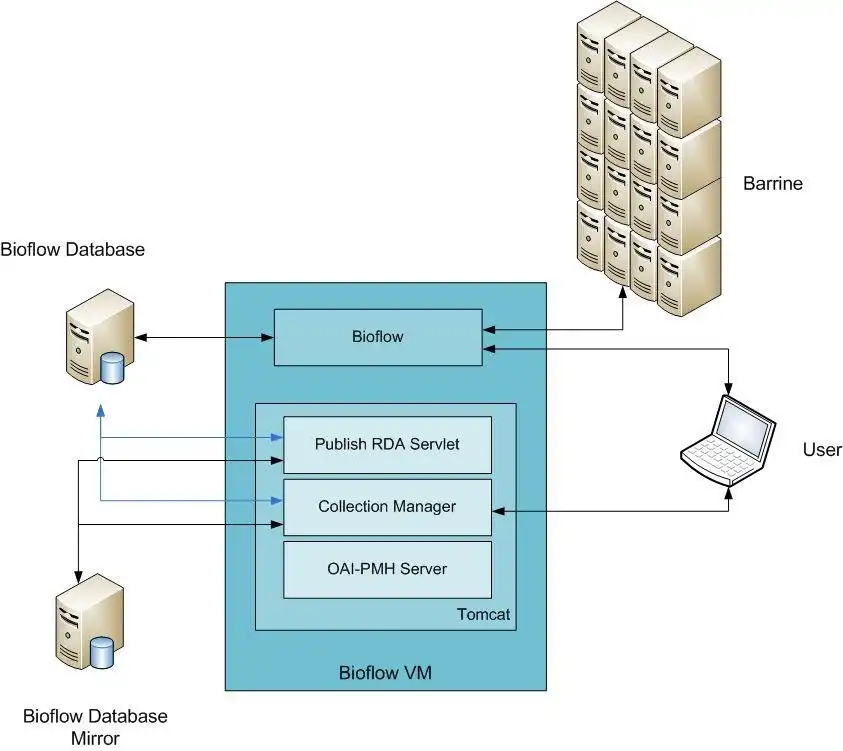
Modern life science research requires bringing together Biology and Information technology. To help make this process easier, the CCG has developed YABI, an Internet based workflow application that is aimed at biologists who wish to conduct bioinformatics analysis. YABI integrates bioinformatics tools and data via an intuitive workflow creation and management environment. YABI can be used to access the NCI-SF in Bioinformatics and the tools available through the Embl Australia EBI mirror hosted at the University of Queensland. With the support of ANDS, QFAB has developed this software system to enable the publication of services and collections based of Yabi workflows and datasets. By sharing the description of their workflows, scientists facilitate the diffusion and re-use of their data. This website provides access to the workflows and datasets that have been made public. By login in, scientist can edit, manage and publish their collection descriptions to the Research Data Australia.
Programming Language
JavaScript, JSP, Java
Database Environment
PostgreSQL (pgsql)
Categories
This is an application that can also be fetched from https://sourceforge.net/projects/yabi-to-rda/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.