This is the Linux app named Alphafold whose latest release can be downloaded as Alphafoldv2.3.2.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named Alphafold with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
Alphafold
DESCRIPTION
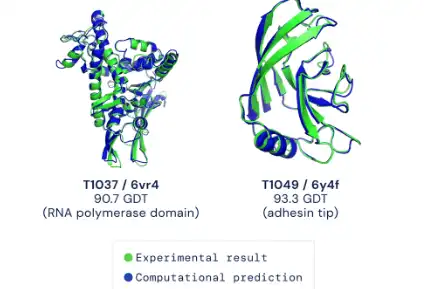
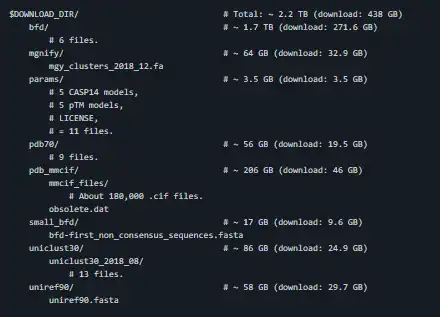
This package provides an implementation of the inference pipeline of AlphaFold v2.0. This is a completely new model that was entered in CASP14 and published in Nature. For simplicity, we refer to this model as AlphaFold throughout the rest of this document. Any publication that discloses findings arising from using this source code or the model parameters should cite the AlphaFold paper. Please also refer to the Supplementary Information for a detailed description of the method. You can use a slightly simplified version of AlphaFold with this Colab notebook or community-supported versions. The total download size for the full databases is around 415 GB and the total size when unzipped is 2.2 TB. Please make sure you have a large enough hard drive space, bandwidth and time to download. We recommend using an SSD for better genetic search performance.
Features
- AlphaFold needs multiple genetic (sequence) databases to run
- While the AlphaFold code is licensed under the Apache 2.0 License, the AlphaFold parameters are made available for non-commercial use
- 5 models which were used during CASP14, and were extensively validated for structure prediction quality
- 5 pTM models, which were fine-tuned to produce pTM
- The simplest way to run AlphaFold is using the provided Docker script
- By default, Alphafold will attempt to use all visible GPU devices
Programming Language
Python
Categories
This is an application that can also be fetched from https://sourceforge.net/projects/alphafold.mirror/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.