This is the Linux app named Automatic cell lineage reconstruction to run in Linux online whose latest release can be downloaded as testData.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named Automatic cell lineage reconstruction to run in Linux online with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
Automatic cell lineage reconstruction to run in Linux online
DESCRIPTION
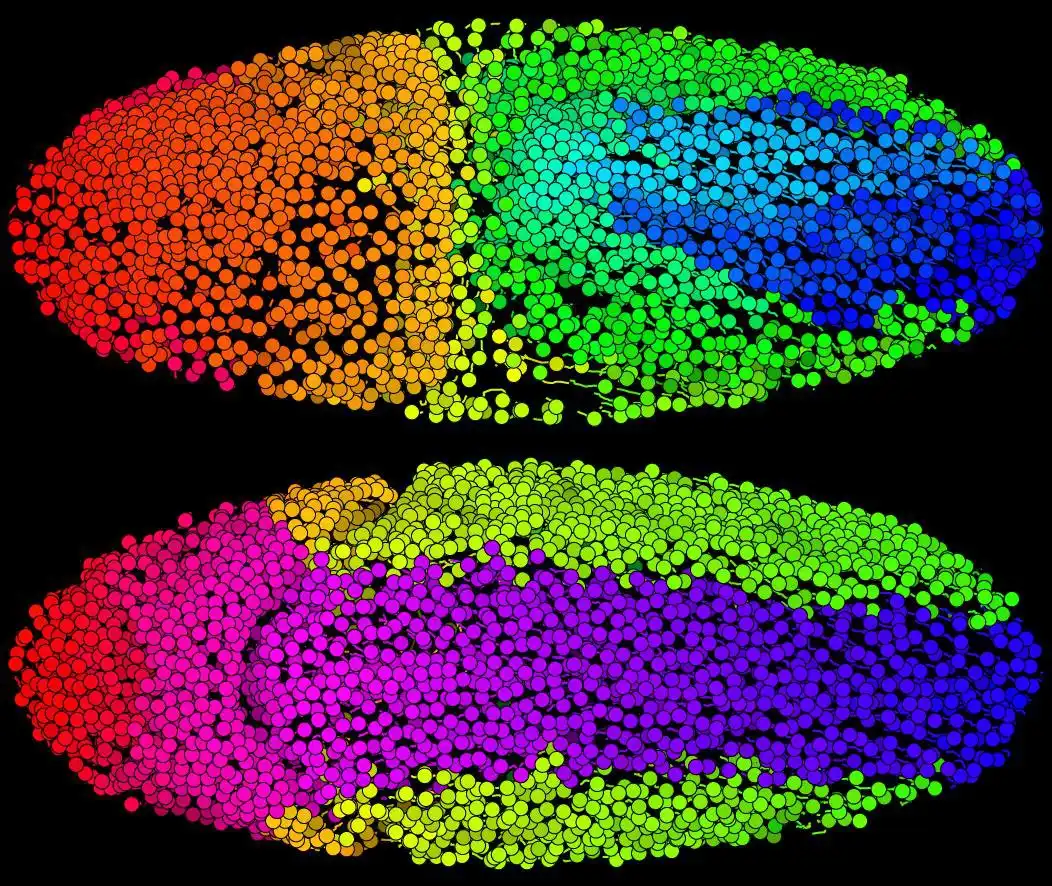
From Amat et al., Nature Methods, 2014*:"The comprehensive reconstruction of cell lineages in complex multicellular organisms is a central goal of developmental biology. We present an open-source computational framework for segmentation and tracking of cell nuclei with high accuracy and speed. We demonstrate its (1) generality, by reconstructing cell lineages in four-dimensional, terabyte-sized image data of fruit-fly, zebrafish and mouse embryos, acquired with three different types of fluorescence microscopes, (2) scalability, by analyzing advanced stages of development with up to 20,000 cells per time point, at 26,000 cells min-1 on a single computer workstation, and (3) ease of use, by adjusting only two parameters across all data sets and providing visualization and editing tools for efficient data curation. Our approach achieves on average 97.0% linkage accuracy across all species and imaging modalities."
*Please cite this paper if you use this code for your research
Audience
Science/Research, End Users/Desktop
User interface
Console/Terminal
Programming Language
C++, C
This is an application that can also be fetched from https://sourceforge.net/projects/tgmm/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.