This is the Linux app named MITP to run in Linux online whose latest release can be downloaded as MITP-1.1.tar.gz. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named MITP to run in Linux online with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
MITP to run in Linux online
DESCRIPTION
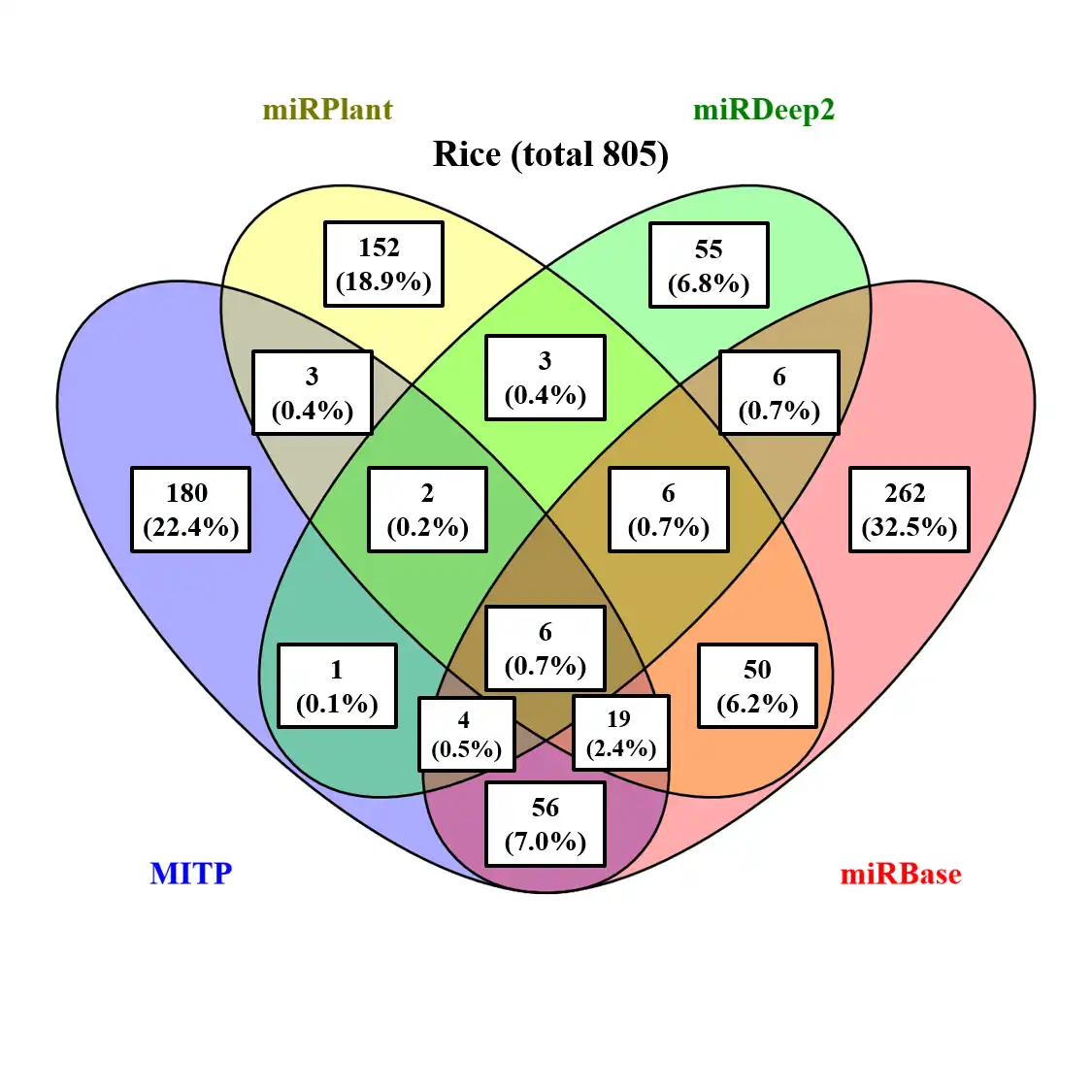
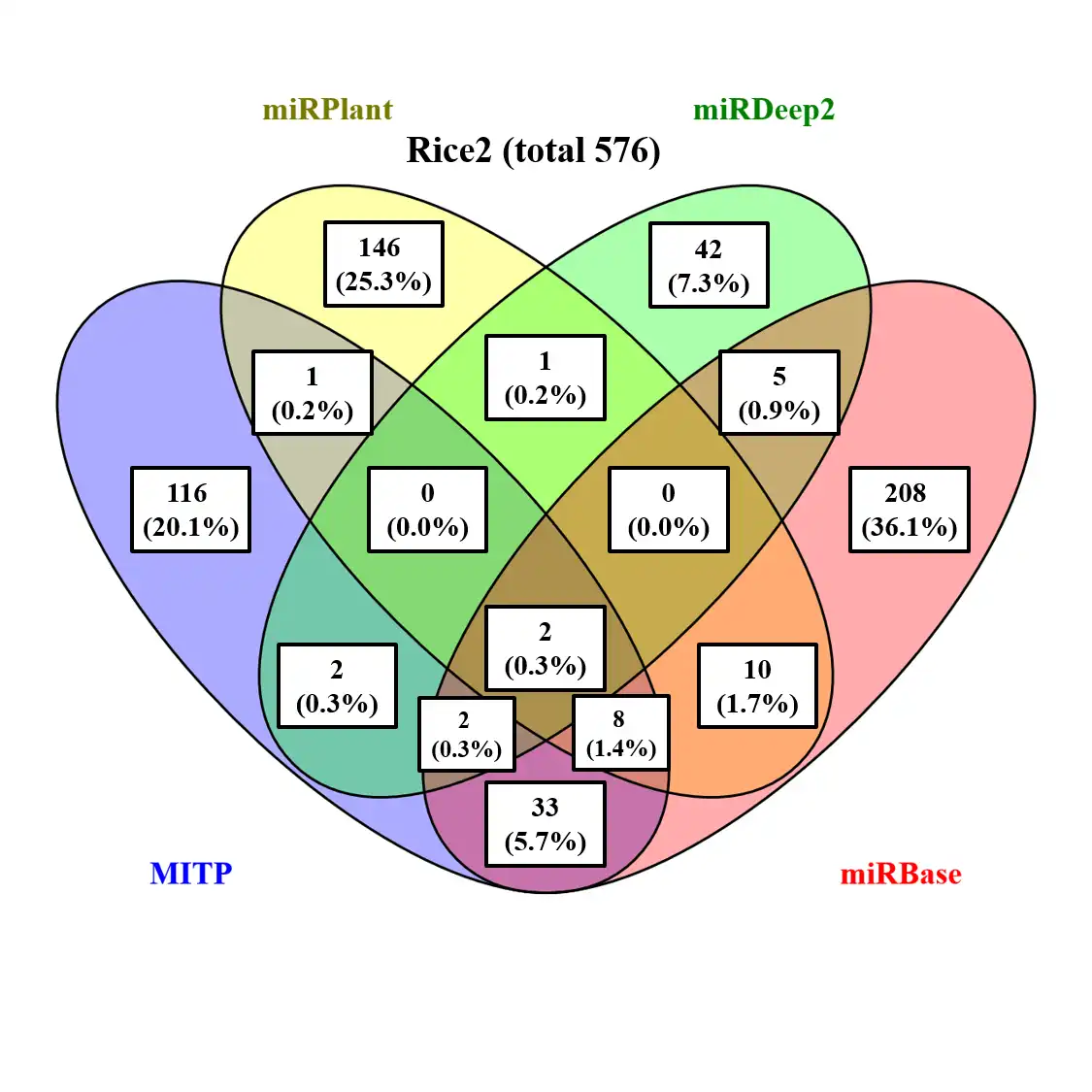
miRNA is a widely known small non-coding RNA which can mediate gene regulation of most important biological processes in plants and animals. Therefore, identification conserve and novel miRNA and their target genes in model and new sequenced species are inevitable. MITP is designed to identify miRNA easily and faster based on sequence mapping result from any mapping software which producing SAM format output result, blast result (default output result) or blat result (default output result). The program provide a step praramter (8 steps) which can allow running program from any step and finishing all remaining steps. You also can run step by step using each step program. Please run these step programs at the same directory for running main program MITP.pl. Some steps are optional (read filter, expression, miRNA class and target). When you select the related parameters belong to these optional steps, the program will run these steps. Otherwise, it skip these steps.Features
- Identify conserve miRNA according to the BLAST result between known miRNA and genome sequence
- Identify novel miRNA based on alignment result in default BLAST output, BLAT output or SAM format file
- Identify samll RNA regions
- Predict miRNA target genes
- Compare miRNA in different species
Audience
Science/Research
Programming Language
Perl
This is an application that can also be fetched from https://sourceforge.net/projects/mitp/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.