This is the Linux app named RNASeq-MATS to run in Linux online whose latest release can be downloaded as testData.tgz. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named RNASeq-MATS to run in Linux online with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
RNASeq-MATS to run in Linux online
DESCRIPTION
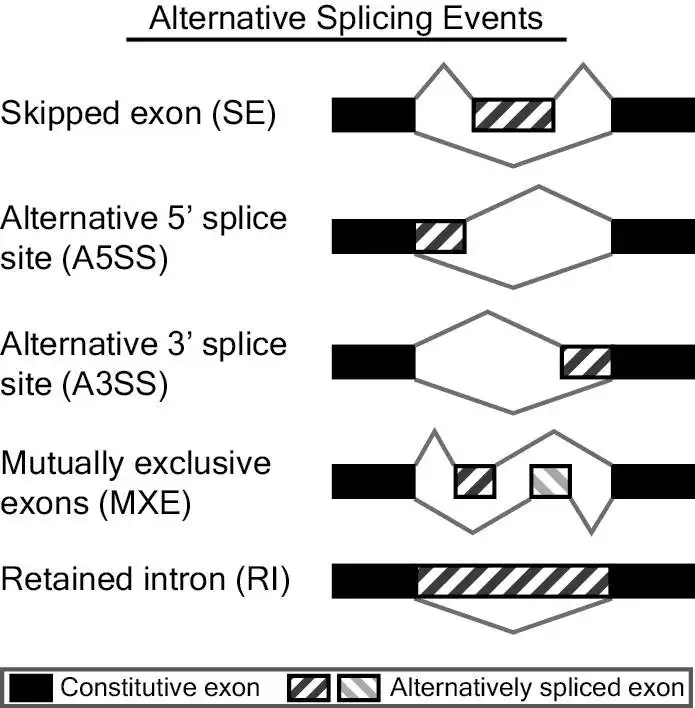
MATS is a computational tool to detect differential alternative splicing events from RNA-Seq data. The statistical model of MATS calculates the P-value and false discovery rate that the difference in the isoform ratio of a gene between two conditions exceeds a given user-defined threshold. From the RNA-Seq data, MATS can automatically detect and analyze alternative splicing events corresponding to all major types of alternative splicing patterns. MATS handles replicate RNA-Seq data from both paired and unpaired study design. More information can be found at http://rnaseq-mats.sourceforge.net.This is an application that can also be fetched from https://sourceforge.net/projects/rnaseq-mats/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.