This is the Linux app named Visualization of Protein-Ligand Graphs whose latest release can be downloaded as vplg_2016_07_18.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named Visualization of Protein-Ligand Graphs with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
Visualization of Protein-Ligand Graphs
DESCRIPTION
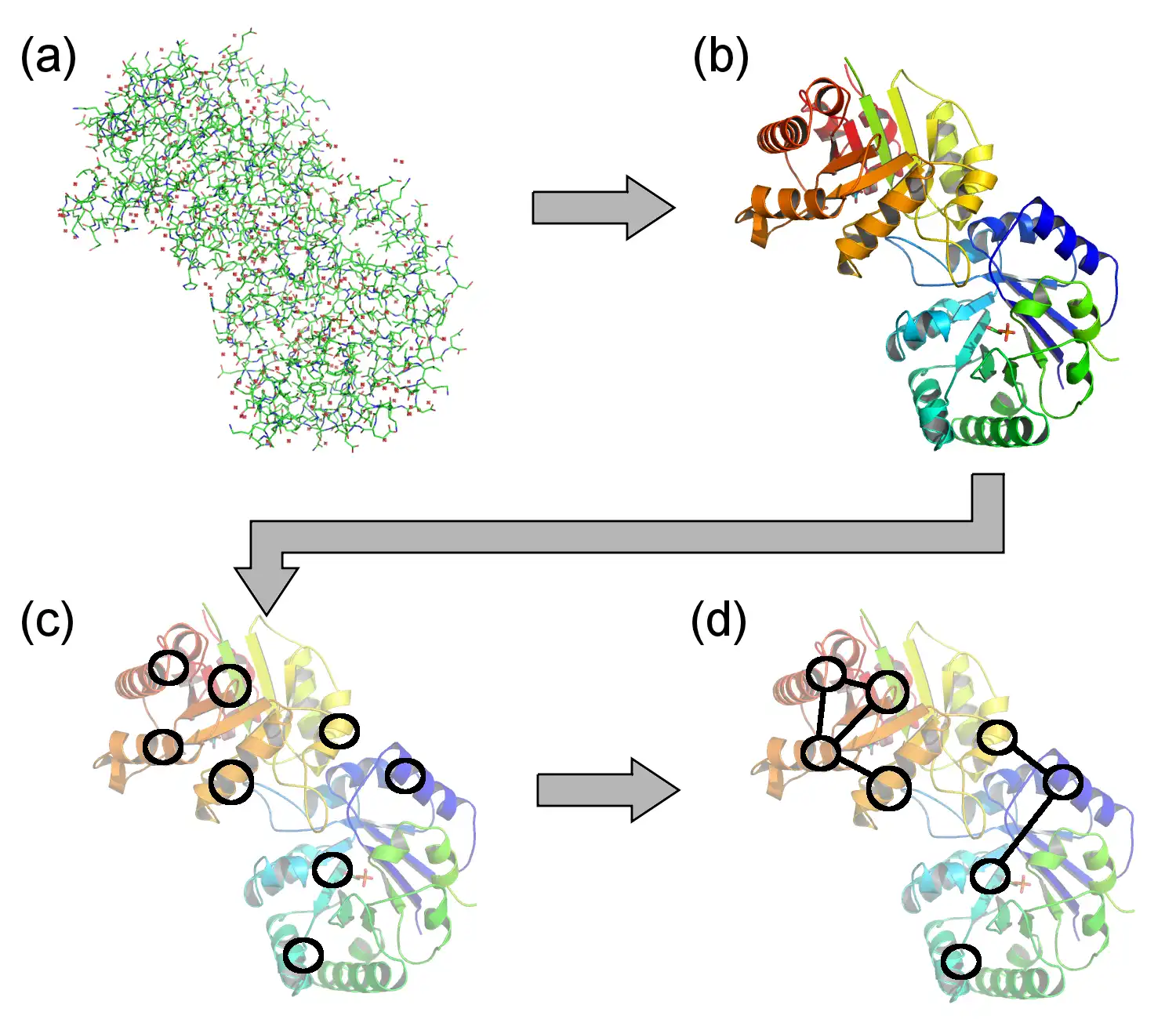
The Visualization of Protein-Ligand Graphs (VPLG) software package computes and visualizes protein graphs. It works on the super-secondary structure level and uses the atom coordinates from PDB files and the SSE assignments of the DSSP algorithm.
VPLG is command line software. If you do not like typing commands, try our PTGL web server: http://ptgl.uni-frankfurt.de/
Features
- Reads 3-dimensional atom data from PDB files and secondary structure assignments from DSSP files
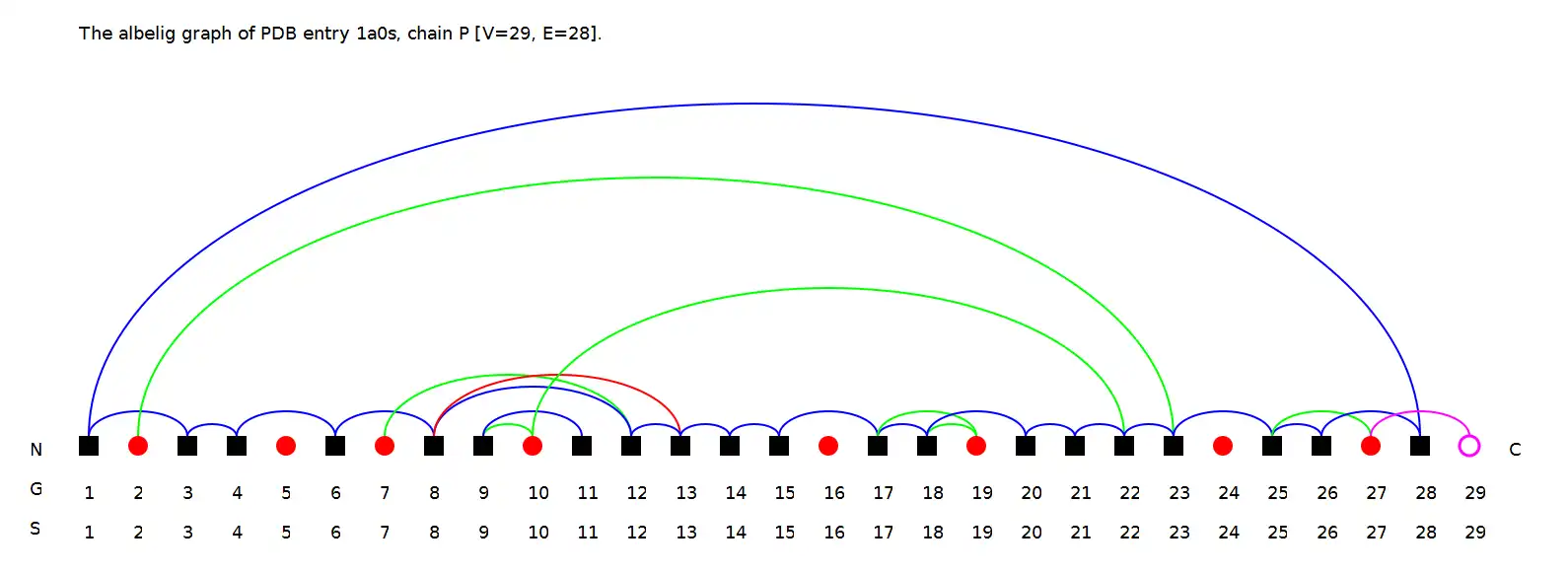
- Computes a protein-ligand graph from the data and visualizes the graph
- Supports output of the graph images in bitmap (.png) and vector (.svg) formats
- Exports protein-ligand graphs in a plain text file in its own format (.plg) that is easy to edit, parse and generate with a computer program as well as in a number of standard graph file formats
- Can read and directly visualize protein graphs from .plg-files
- Free Open Source Software (FOSS)
- Database support (completely optional, no database server required by default)
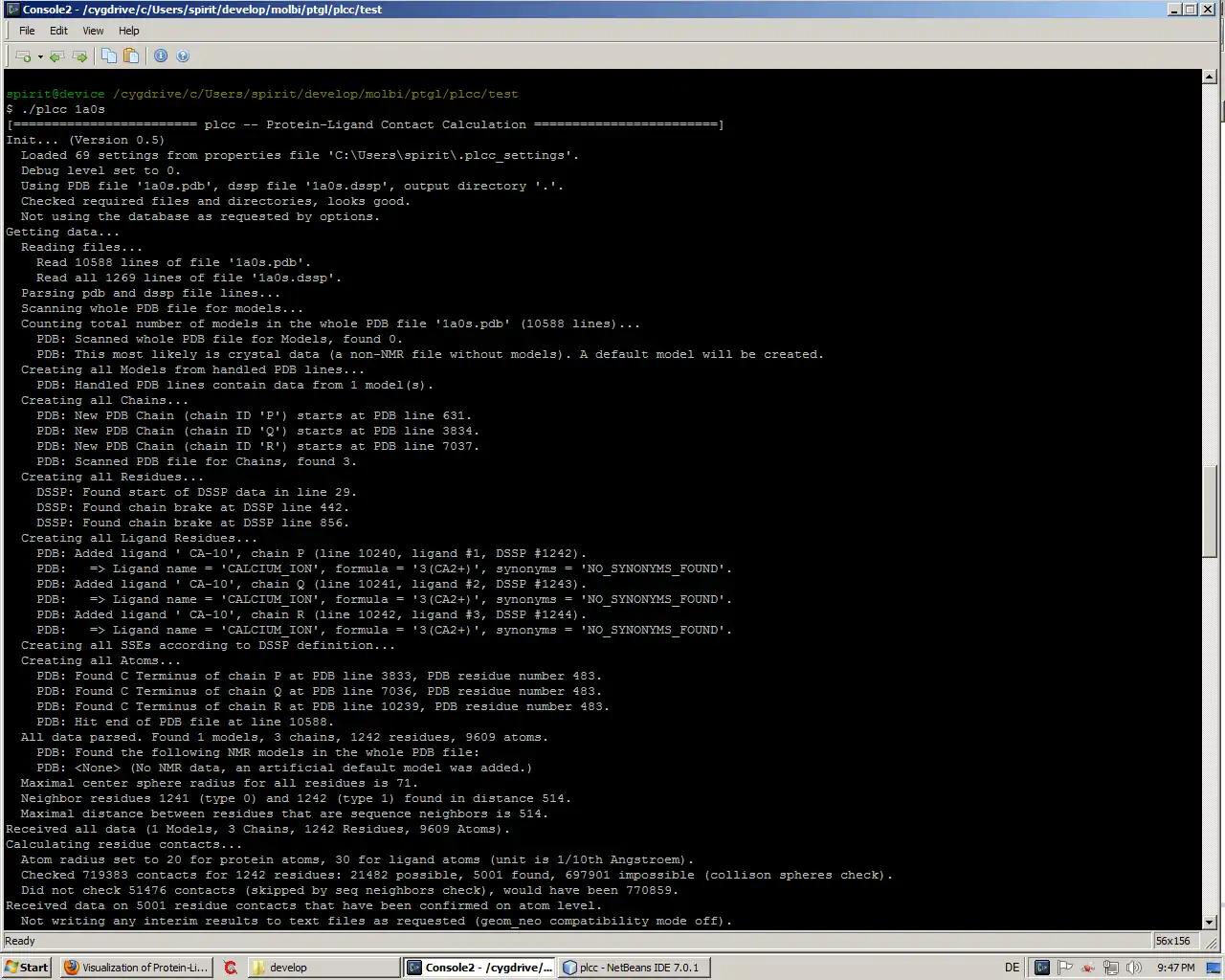
- Command line interface allows for easy batch processing via shell scripts
- Allows filtering of ligands by atom number
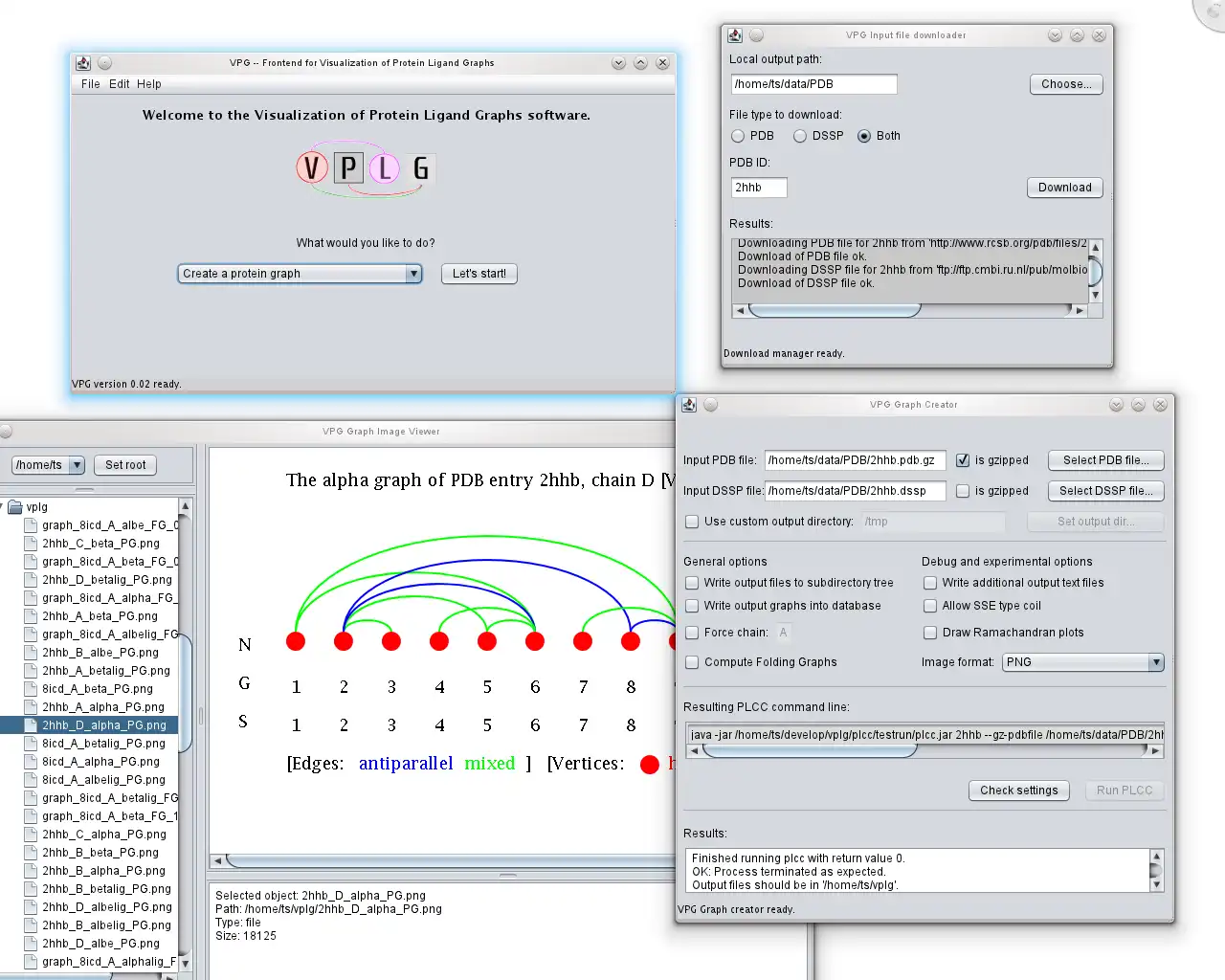
- New GUI written in Java, no need to type console commands anymore!
- The software that powers the PTGL web server: http://ptgl.uni-frankfurt.de/
Audience
Science/Research, Education, Advanced End Users
User interface
Java Swing, Console/Terminal, Command-line
Programming Language
Java
Database Environment
JDBC
Categories
This is an application that can also be fetched from https://sourceforge.net/projects/vplg/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.