This is the Windows app named Covidex whose latest release can be downloaded as Covidex_2021_11_27.rar. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named Covidex with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start any OS OnWorks online emulator from this website, but better Windows online emulator.
- 5. From the OnWorks Windows OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application and install it.
- 7. Download Wine from your Linux distributions software repositories. Once installed, you can then double-click the app to run them with Wine. You can also try PlayOnLinux, a fancy interface over Wine that will help you install popular Windows programs and games.
Wine is a way to run Windows software on Linux, but with no Windows required. Wine is an open-source Windows compatibility layer that can run Windows programs directly on any Linux desktop. Essentially, Wine is trying to re-implement enough of Windows from scratch so that it can run all those Windows applications without actually needing Windows.
SCREENSHOTS
Ad
Covidex
DESCRIPTION
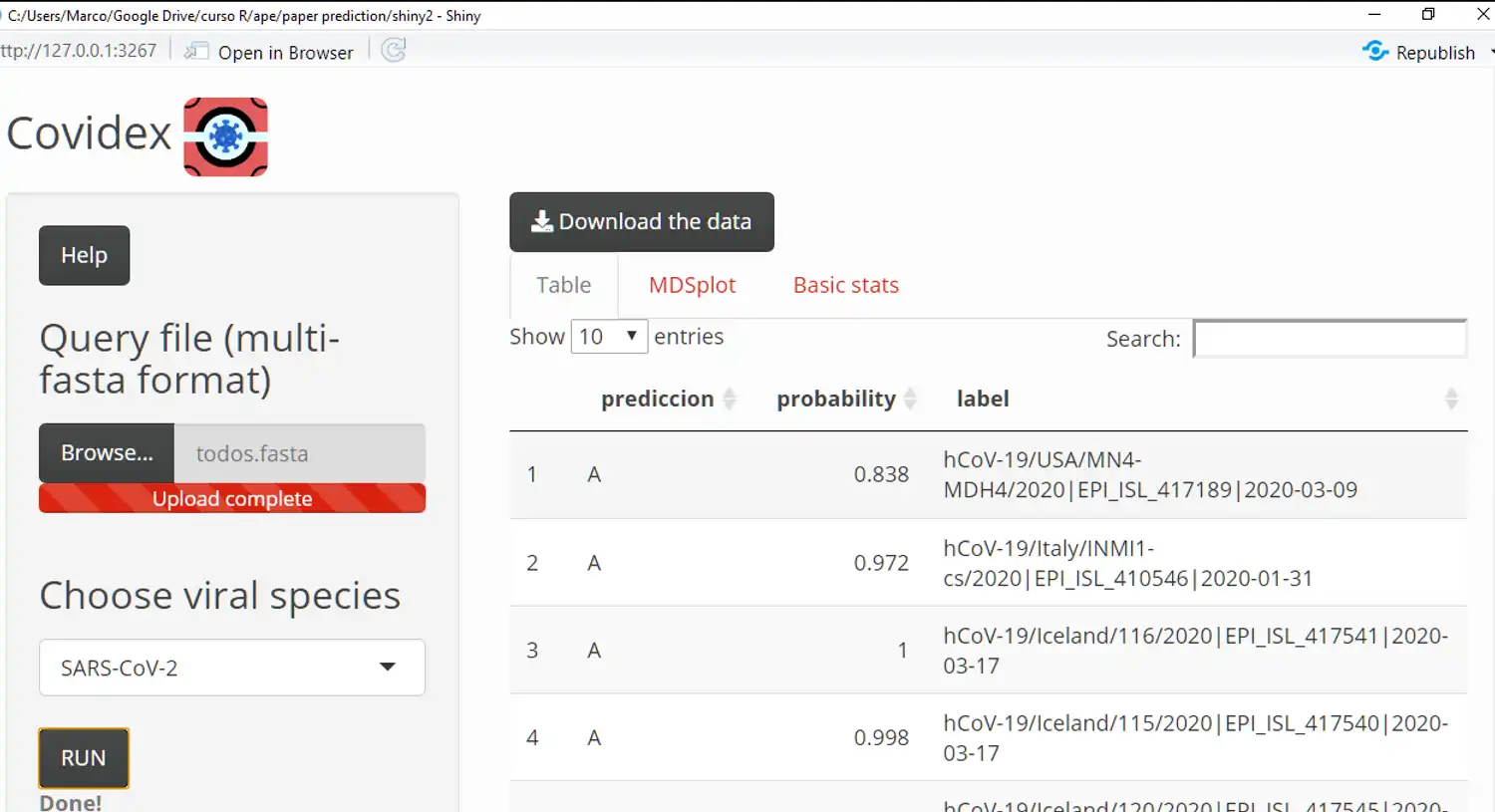
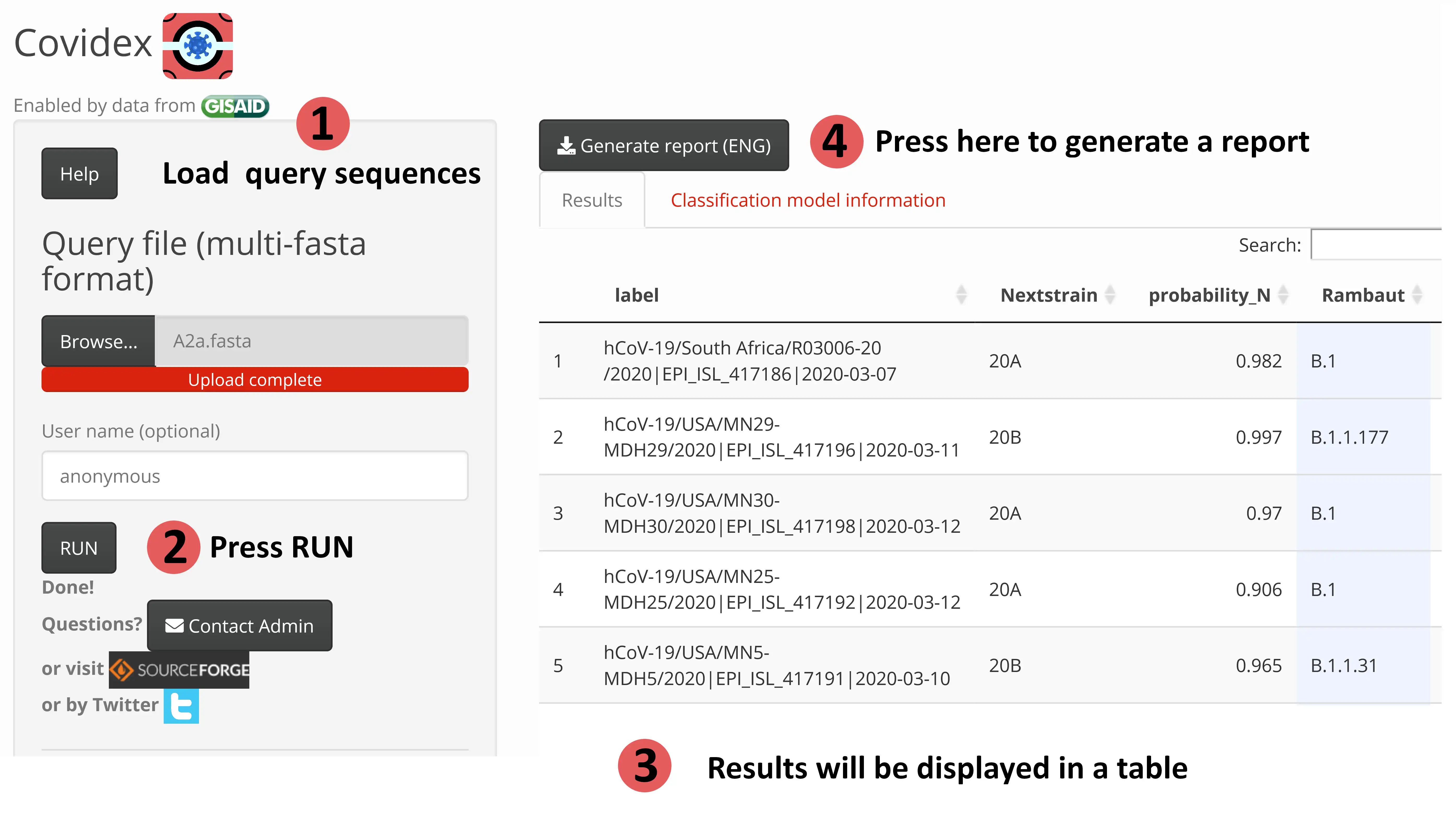
Viral subtypes or clades represent clusters among isolates from the global population of a defined species. Subtypification is relevant for studies on virus epidemiology, evolution and pathogenesis. In this sense, Covidex was developed as an open source alignment-free machine learning subtyping tool. It is a shiny app that allows fast and accurate classification of viral genomes in pre-defined clusters. If more than 1000 sequences are loaded the tool will run in multithread mode. Capable of classifying 16000 genome sequences in less than a minute (AMD Ryzen 7 1700 8-core Processor 3 GHz)
For a Web-based version of the app (only for small datasets: 100 seqs max) please go to
http://covidex.unlu.edu.ar
If you use Covidex please consider citing the following preprint:
https://biorxiv.org/cgi/content/short/2020.08.21.261347v1
If you think my work is useful you can buy me a coffee! https://www.buymeacoffee.com/mcacciabue
Features
- Subtyping of sequences
- User-friendly interface
- Summary plots
- Portable version for Windows (No installation needed)
- Multithread mode
Audience
Science/Research, End Users/Desktop
Programming Language
R
This is an application that can also be fetched from https://sourceforge.net/projects/covidex/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.