This is the Windows app named Javamony whose latest release can be downloaded as Javamony.jar. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named Javamony with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start any OS OnWorks online emulator from this website, but better Windows online emulator.
- 5. From the OnWorks Windows OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application and install it.
- 7. Download Wine from your Linux distributions software repositories. Once installed, you can then double-click the app to run them with Wine. You can also try PlayOnLinux, a fancy interface over Wine that will help you install popular Windows programs and games.
Wine is a way to run Windows software on Linux, but with no Windows required. Wine is an open-source Windows compatibility layer that can run Windows programs directly on any Linux desktop. Essentially, Wine is trying to re-implement enough of Windows from scratch so that it can run all those Windows applications without actually needing Windows.
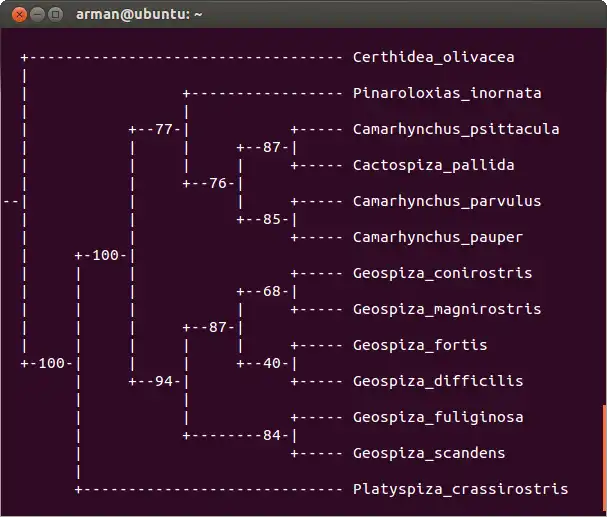
SCREENSHOTS
Ad
Javamony
DESCRIPTION
Based on the not-so-successful Pysimony (https://sourceforge.net/projects/pysimony/), the same determined student takes another go at the phylogenetic problem.
Javamony is invoked as follows:
java -jar Javamony.jar [input.fasta] [random / stepwise (starting tree)] [# of bootstraps] [outgroup taxon #1] [outgroup taxon #2] ...
Not meant as a competitive phylogenetic inference program, Javamony is an opportunity for me to acquire the Java language while learning to address and solve fundamental problems in phylogenetics. Therefore, for my own educational benefit, all code is original. Of course, there are probably a good deal of mistakes as well.
I distribute Javamony, as I did Pysimony, hoping that it will be of educational value to someone else or at least vaguely amusing.
Upcoming features will be:
- Support for Amino Acid sequences
- Support for additional file formats (e.g. Nexus)
- Multithreading
- Additional scoring methods (e.g. maximum likelihood)
Features
- Uses maximum parsimony
- Reads a FASTA input file (DNA only)
- User-established outgroup
- Random and stepwise-addition starting trees
- Uses SPR (subtree pruning and regrafting) tree search
- Performs bootstrapping
- Outputs standard Newick format with bootstrap support and branchlengths
- Draws final tree with bootstrap support
Audience
Science/Research, Education
User interface
Command-line
Programming Language
Java
Categories
This is an application that can also be fetched from https://sourceforge.net/projects/javamony/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.