This is the Windows app named NGSEP whose latest release can be downloaded as NGSEPwindows_4.1.0.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named NGSEP with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start any OS OnWorks online emulator from this website, but better Windows online emulator.
- 5. From the OnWorks Windows OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application and install it.
- 7. Download Wine from your Linux distributions software repositories. Once installed, you can then double-click the app to run them with Wine. You can also try PlayOnLinux, a fancy interface over Wine that will help you install popular Windows programs and games.
Wine is a way to run Windows software on Linux, but with no Windows required. Wine is an open-source Windows compatibility layer that can run Windows programs directly on any Linux desktop. Essentially, Wine is trying to re-implement enough of Windows from scratch so that it can run all those Windows applications without actually needing Windows.
SCREENSHOTS
Ad
NGSEP
DESCRIPTION
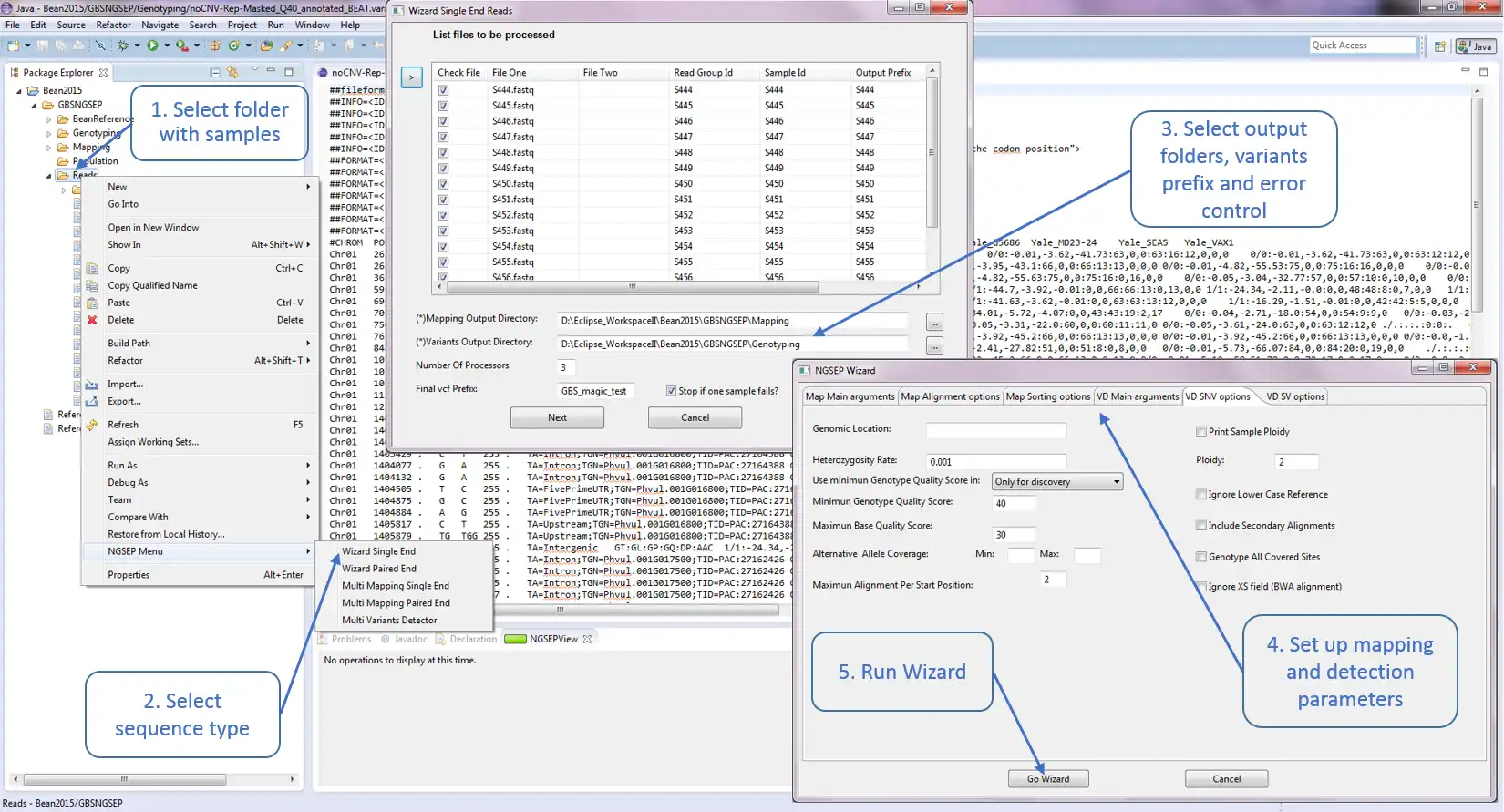
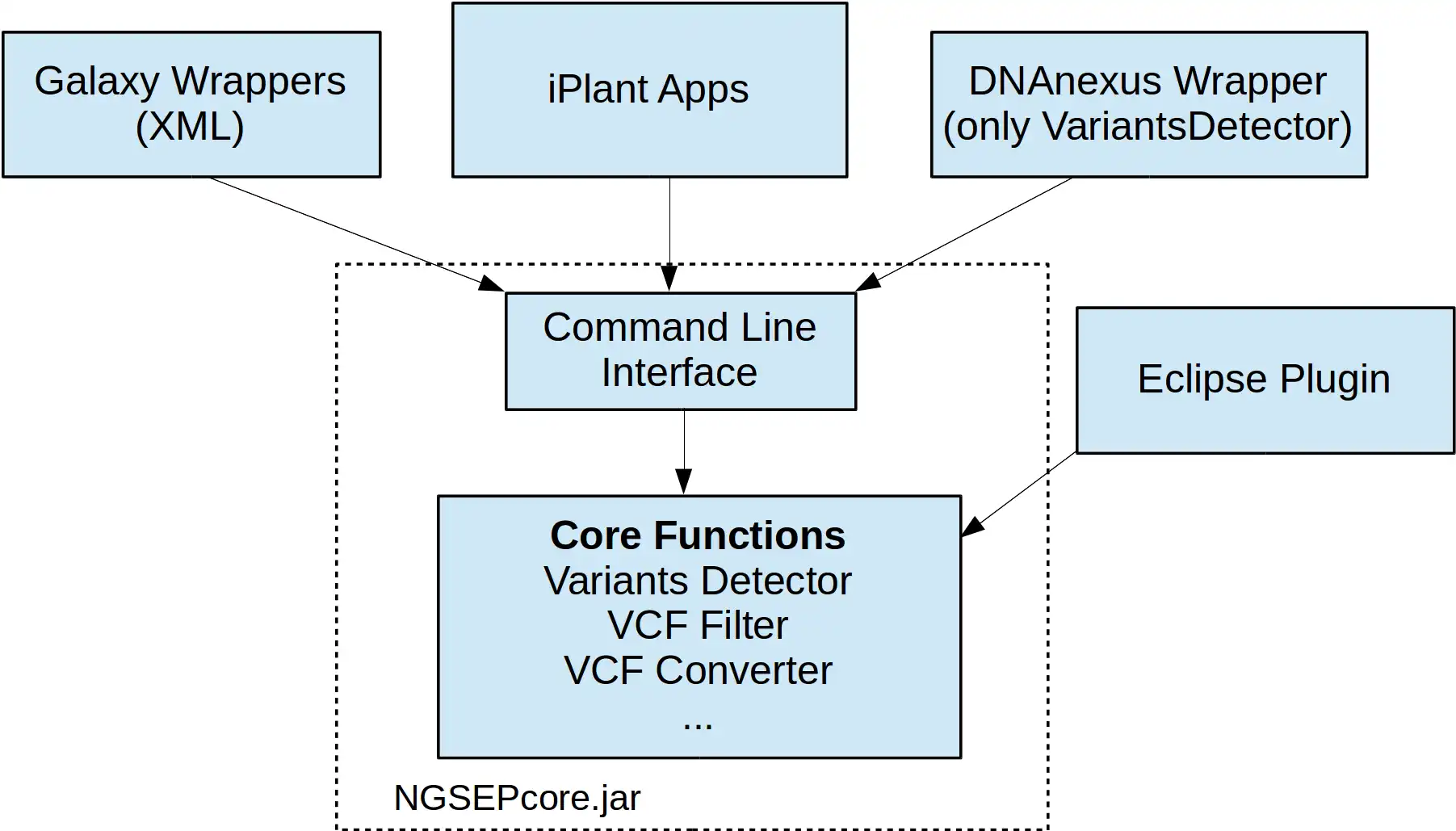
NGSEP is an integrated framework for analysis of DNA high throughput sequencing data. The main use of NGSEP is the construction and downstream analysis of large datasets of genomic variation. NGSEP performs accurate detection and genotyping of Single Nucleotide Variants (SNVs), small and large indels, short tandem repeats (STRs), inversions, and Copy Number Variants (CNVs). NGSEP also provides modules for functional annotation, filtering, format conversion, comparison, clustering, imputation, introgression analysis and different kinds of statistics. A complete list of functionalities is available in our wiki (https://sourceforge.net/p/ngsep/wiki/Home/).
NEWS: We just made available the first version of our de-novo genomes assembler. Further details are available in the wiki.
Features
- De-novo genomes assembly
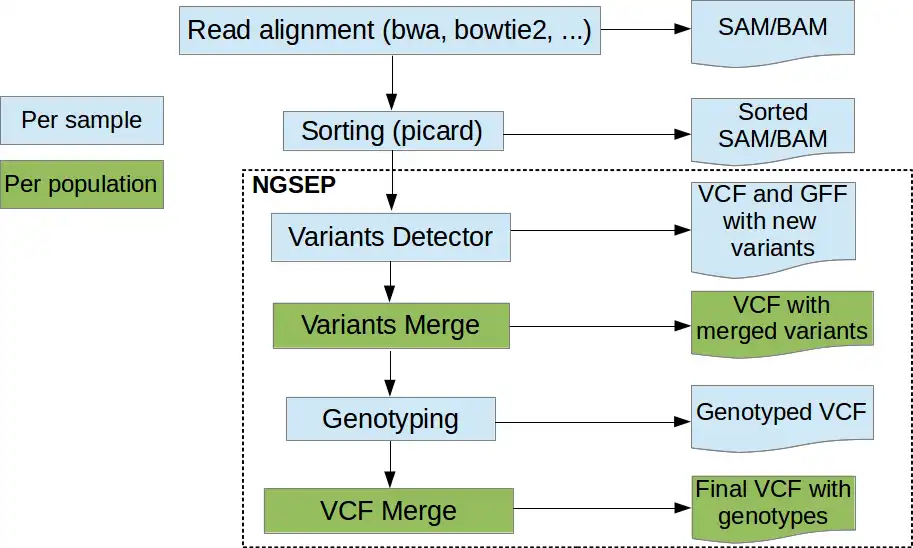
- Alignment of raw reads to a reference genome
- SNPs, CNVs and Structural Variants detection
- VCF manipulation: functional annotation, merge, filter, compare, format conversion, imputation
- Reads Demultiplexing
- Alignment of annotated genome assemblies
- Statistics and filtering on transcriptome annptations in GFF3 format
Audience
Healthcare Industry, Science/Research, Other Audience, Agriculture
User interface
Java SWT, Console/Terminal, Eclipse
Programming Language
Java
This is an application that can also be fetched from https://sourceforge.net/projects/ngsep/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.