This is the Windows app named RNA@ whose latest release can be downloaded as RNAAT-V1.1.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named RNA@ with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start any OS OnWorks online emulator from this website, but better Windows online emulator.
- 5. From the OnWorks Windows OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application and install it.
- 7. Download Wine from your Linux distributions software repositories. Once installed, you can then double-click the app to run them with Wine. You can also try PlayOnLinux, a fancy interface over Wine that will help you install popular Windows programs and games.
Wine is a way to run Windows software on Linux, but with no Windows required. Wine is an open-source Windows compatibility layer that can run Windows programs directly on any Linux desktop. Essentially, Wine is trying to re-implement enough of Windows from scratch so that it can run all those Windows applications without actually needing Windows.
SCREENSHOTS
Ad
RNA@
DESCRIPTION
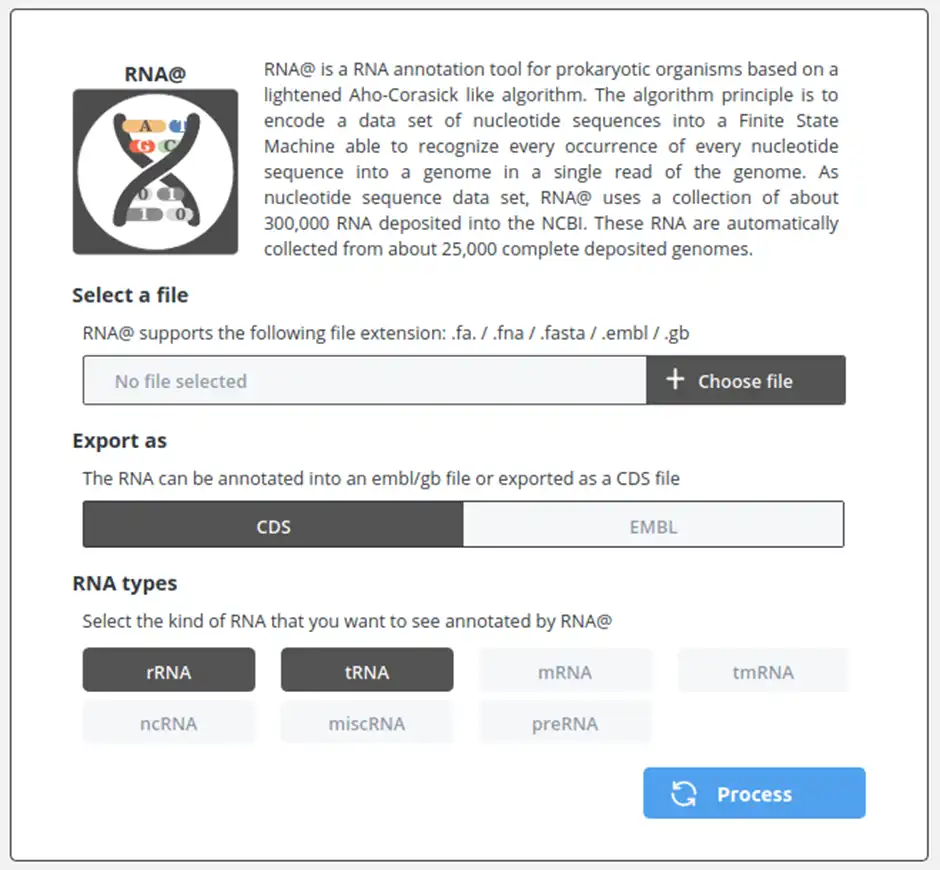
RNA@ is a RNA annotation tool for prokaryotic organisms based on a lightened Aho-Corasick like algorithm. The algorithm principle is to encode a data set of nucleotide sequences into a Finite State Machine able to recognize every occurrence of every nucleotide sequence into a genome in a single read of the genome. As nucleotide sequence data set, RNA@ uses a collection of about 300,000 RNA deposited into the NCBI. These RNA are automatically collected from about 25,000 complete deposited genomes.
You will find at https://drive.google.com/file/d/1lEj5SSHpQSv54nOroxBcS_jvLFRIKNmm/view?usp=sharing" the RNA re-annotated by RNA@ for 30,558 genomes of Archaea and Bacteria (all the archaea and bacteria genomes marked as "Complete Genomes" or "Chromosome" among the genomes deposited in the NCBI day June 06 of 2021.
You can also download the last version of the database at http://rnaat.ufpa.br
This is an application that can also be fetched from https://sourceforge.net/projects/rnaat/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.