This is the Windows app named SUPERmerge to run in Windows online over Linux online whose latest release can be downloaded as SUPERmerge.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named SUPERmerge to run in Windows online over Linux online with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start any OS OnWorks online emulator from this website, but better Windows online emulator.
- 5. From the OnWorks Windows OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application and install it.
- 7. Download Wine from your Linux distributions software repositories. Once installed, you can then double-click the app to run them with Wine. You can also try PlayOnLinux, a fancy interface over Wine that will help you install popular Windows programs and games.
Wine is a way to run Windows software on Linux, but with no Windows required. Wine is an open-source Windows compatibility layer that can run Windows programs directly on any Linux desktop. Essentially, Wine is trying to re-implement enough of Windows from scratch so that it can run all those Windows applications without actually needing Windows.
SCREENSHOTS
Ad
SUPERmerge to run in Windows online over Linux online
DESCRIPTION
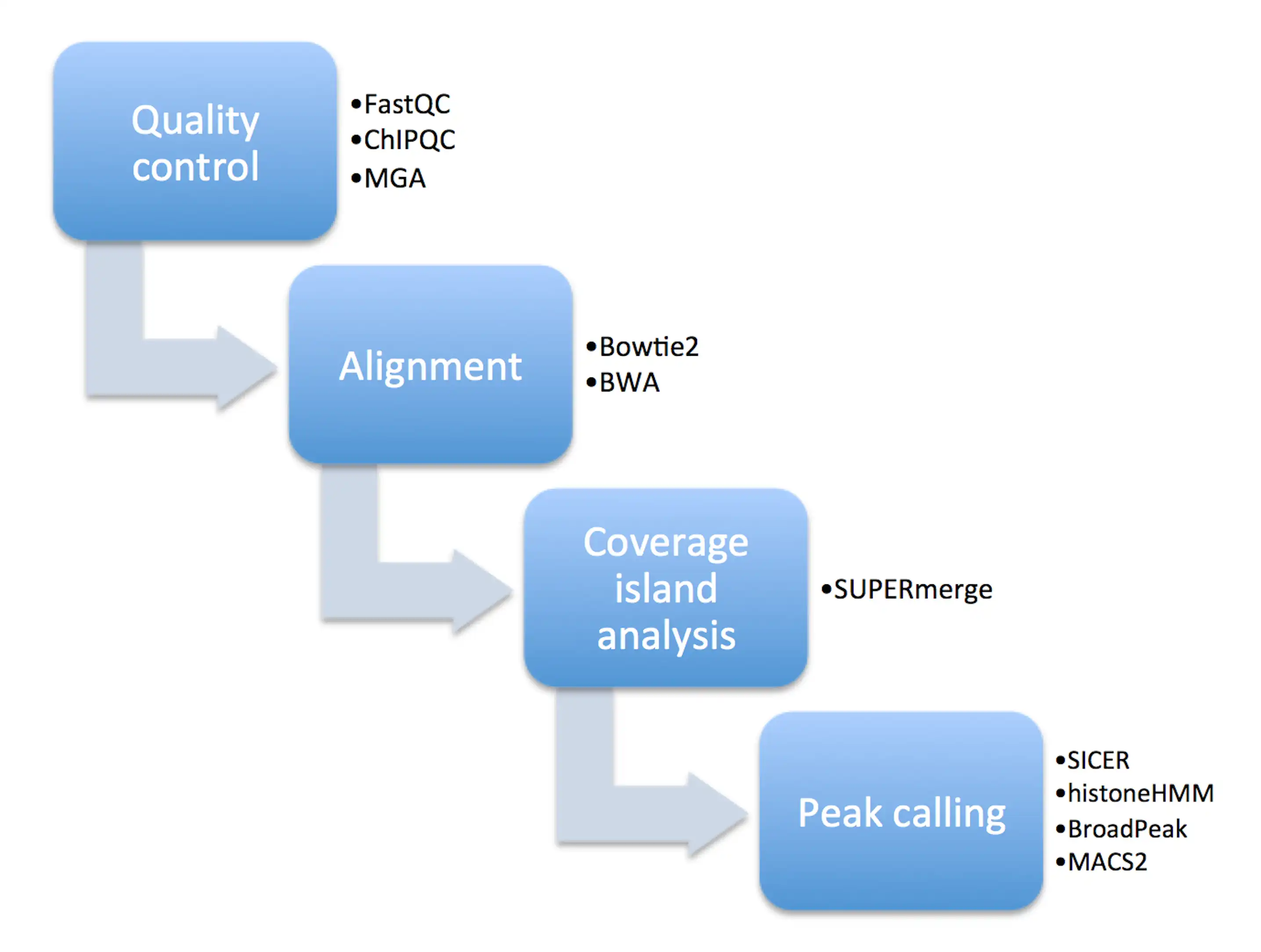
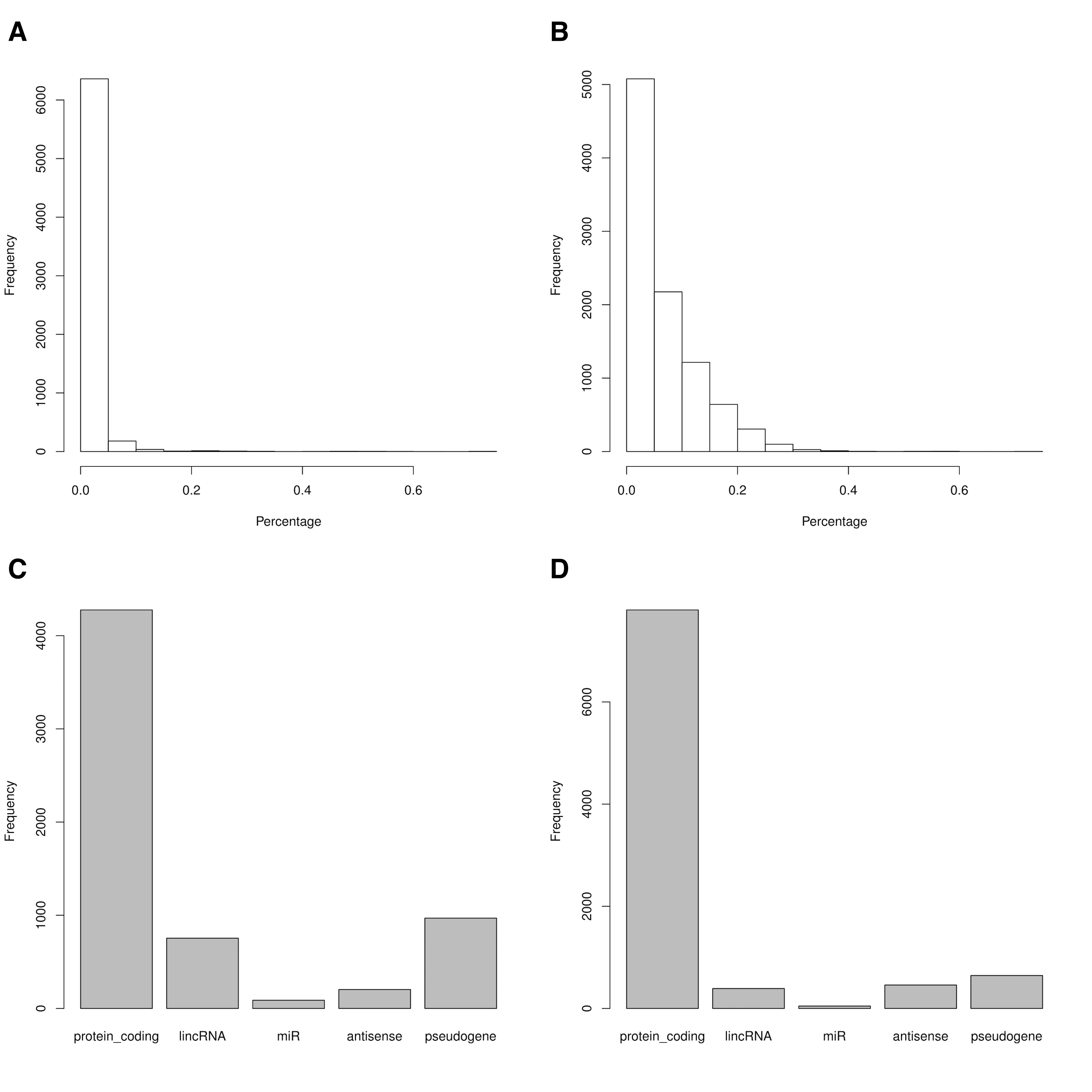
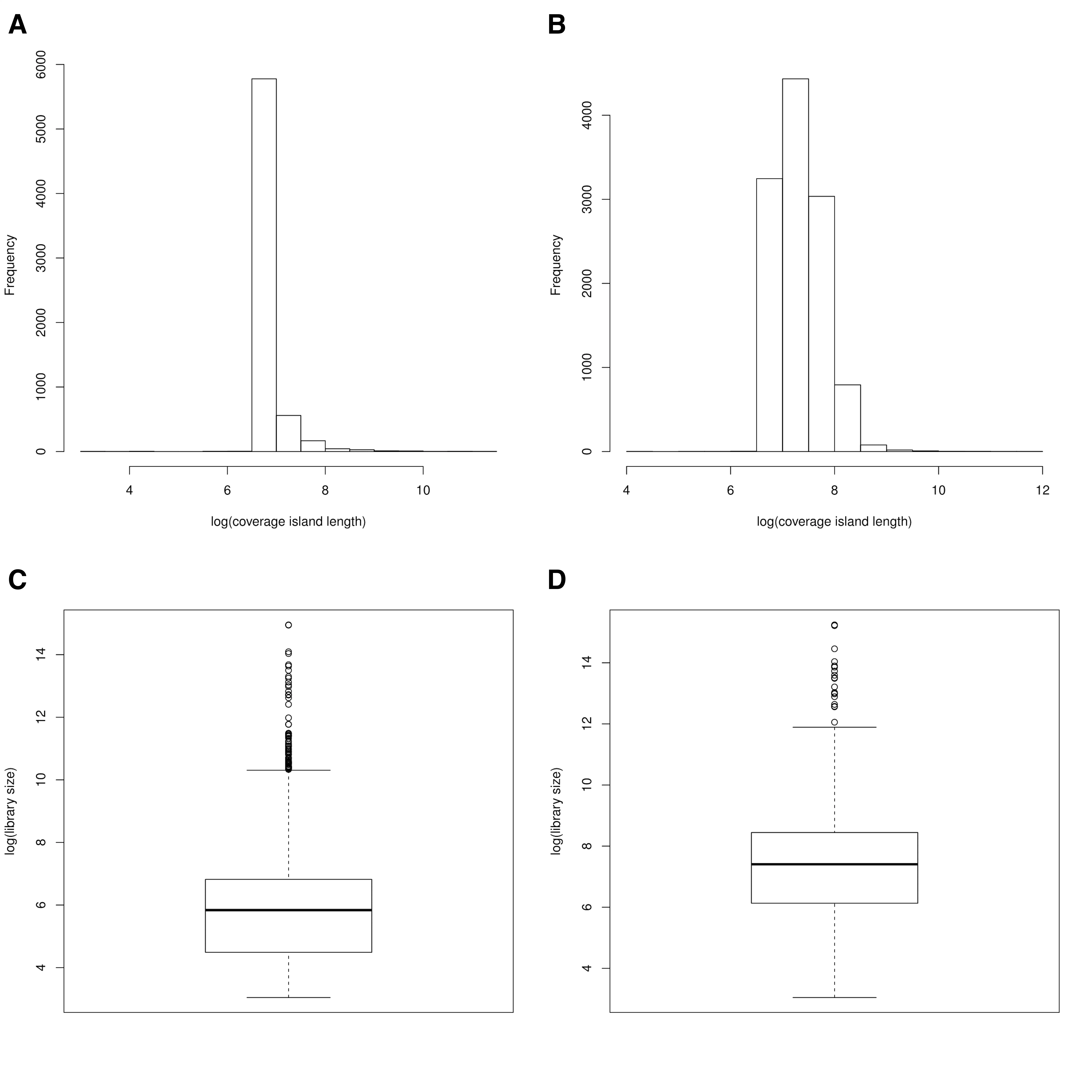
SUPERmerge is a ChIP-seq read pileup analysis and annotation algorithm for investigating alignment (BAM) files of diffuse histone modification ChIP-seq datasets with broad chromatin domains at a single base pair resolution level. SUPERmerge allows flexible regulation of a variety of read pileup parameters, thereby revealing how read islands aggregate into areas of coverage across the genome and what annotation features they map to within individual biological replicates. SUPERmerge is especially useful for investigating low sample size ChIP-seq experiments in which epigenetic histone modifications (e.g., H3K9me1, H3K27me3) result in inherently broad peaks with a diffuse range of signal enrichment spanning multiple consecutive genomic loci and annotated features.Features
- ChIP-seq
- epigenetics
- bioinformatics
- computational biology
Audience
Science/Research
Programming Language
C
This is an application that can also be fetched from https://sourceforge.net/projects/supermerge/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.