This is the Linux app named P3BSseq to run in Linux online whose latest release can be downloaded as P3BSseq_v2.1.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named P3BSseq to run in Linux online with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
P3BSseq to run in Linux online
DESCRIPTION
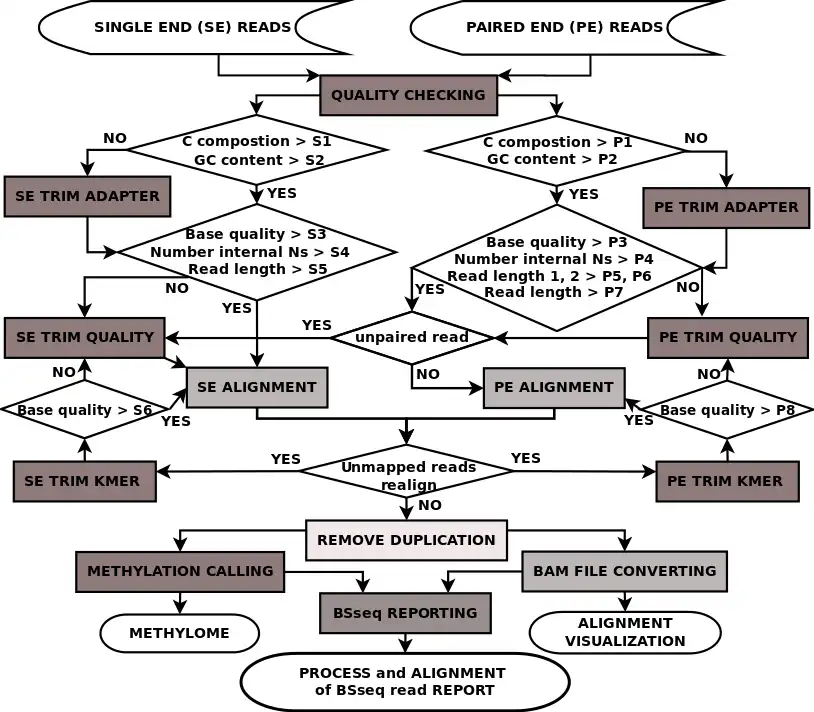
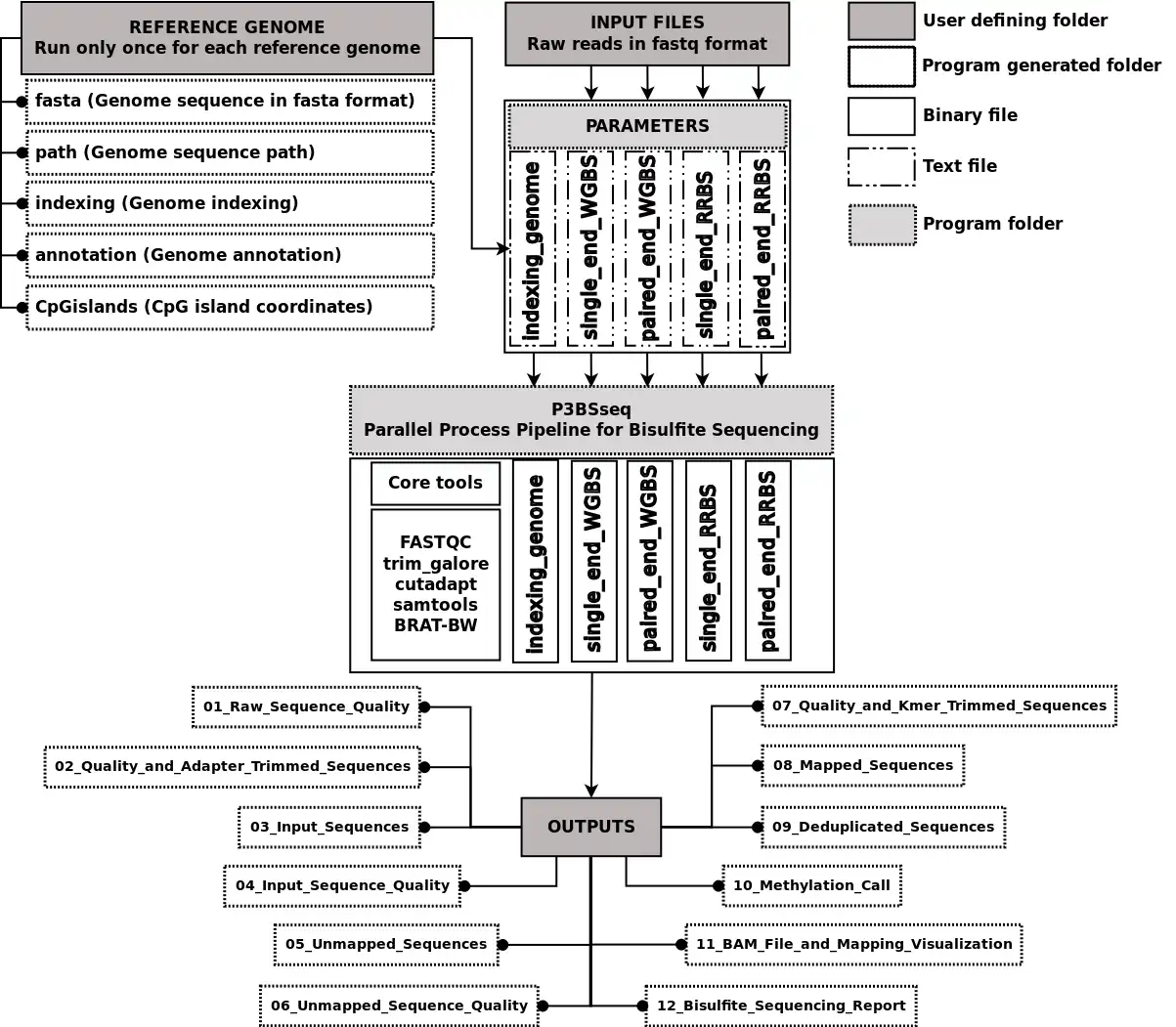
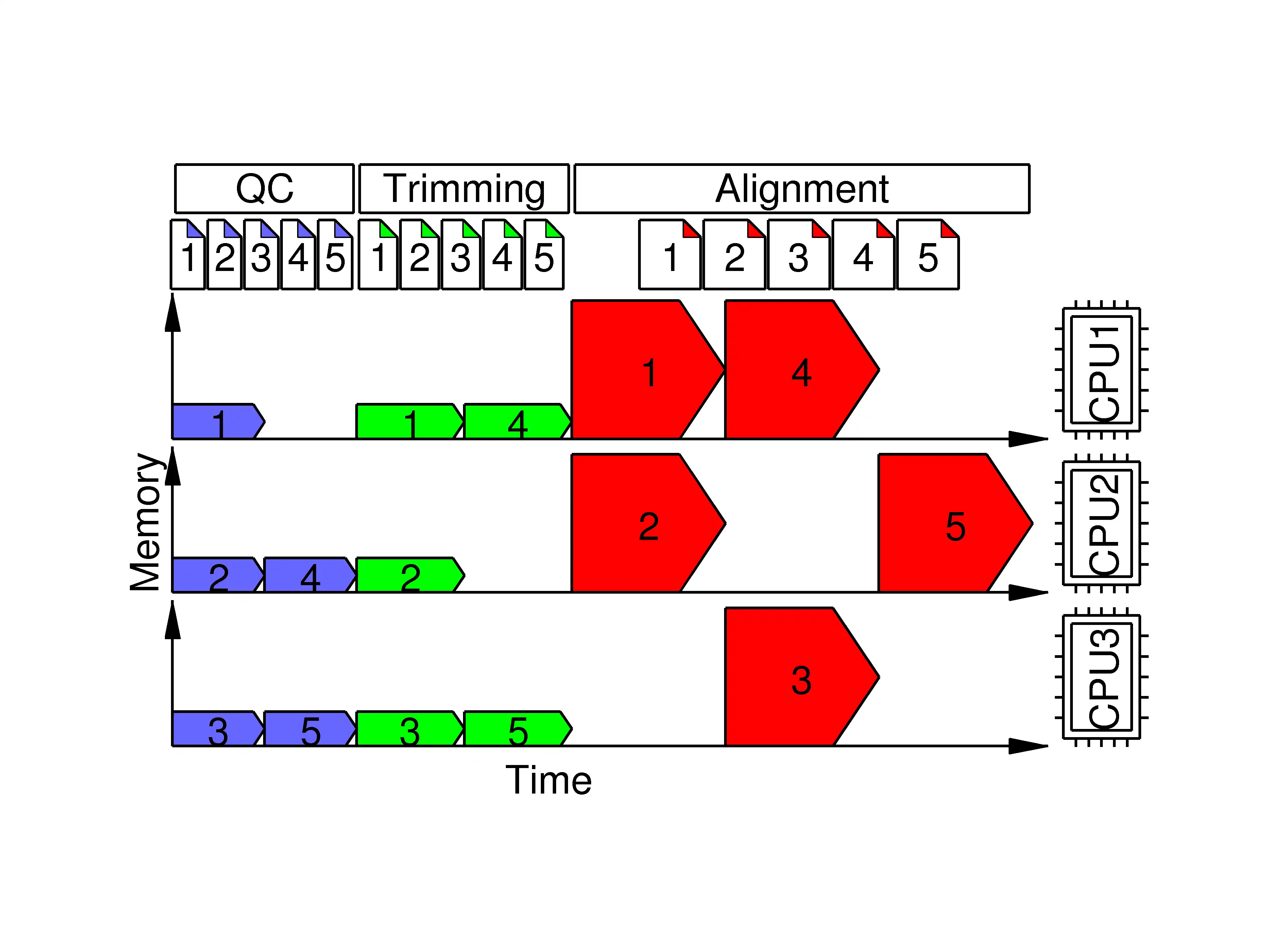
Bisulfite sequencing (BSseq) processing is among the most cumbersome next generation sequencing (NGS) applications. Though some BSseq processing tools are available, they are scattered, require puzzling parameters and are running-time and memory-usage demanding. We have developed P3BSseq, a parallel processing pipeline for fast, accurate and automatic analysis of BSseq reads that trims, aligns, annotates, records the intermediate results, performs bisulfite conversion quality assessment, generates BED methylome and report files following the NIH standards. P3BSseq outperforms the known BSseq mappers regarding running time, computer hardware requirements. We optimized the P3BSseq parameters for both directional and non-directional libraries, and for both single-end and paired-end reads of Whole Genome and Reduced Representation BSseq. P3BSseq is a user-friendly streamlined solution for BSseq upstream analysis, requiring only basic computer and NGS knowledge.Features
- Next Generation Sequencing (NGS) data processing
- DNA methylomics analysis
- Parallel processing
- Reduced Representation of Bisulfite Sequencing (RRBS)
Audience
Science/Research
User interface
Console/Terminal
Programming Language
Python
This is an application that can also be fetched from https://sourceforge.net/projects/p3bsseq/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.