This is the Linux app named Swift Sequence Alignment Program to run in Linux online whose latest release can be downloaded as swift-0.11.2.tgz. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named Swift Sequence Alignment Program to run in Linux online with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
Swift Sequence Alignment Program to run in Linux online
DESCRIPTION
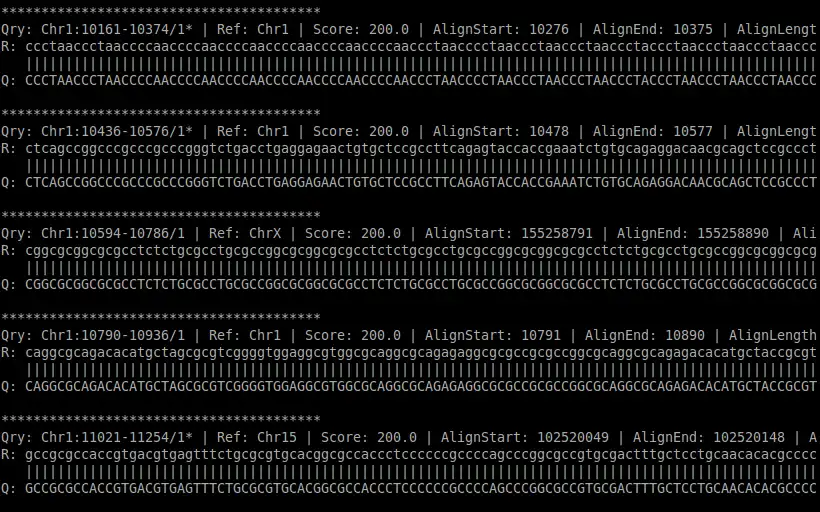
Swift is a DNA sequence alignment program that produces gapped alignment using the Smith-Waterman algorithm. It takes in a query file (FASTA format) and a reference file (FASTA format) as input. It outputs the reference name, read name, gapped alignment, alignment score, alignment start and end positions, and alignment length.I gave a talk on Swift in the GPU Technology Conference 2012. The talk can be accessed at http://www.gputechconf.com/gtcnew/on-demand-GTC.php?sessionTopic=58&searchByKeyword=&submit=&select=+&sessionEvent=&sessionYear=&sessionFormat=#1303.
To install and run Swift, please refer to the Wiki page: http://sourceforge.net/p/swiftseqaligner/wiki/Home/
If you need further help installing or running Swift, please contact Pankaj Gupta at [email protected].
Features
- Produces gapped alignment using Smith-Waterman algorithm
- Smith-Waterman algorithm runs on the GPU (Graphics Processing Unit)
- Accepts a read file (FASTA format) and a reference file (FASTA format) as inputs
- Outputs reference name, read name, gapped alignment, alignment score, alignment start and end positions, and alignment length
- Supports Illumina reads (fixed length)
- Requirements: Linux OS; 4 GB system memory; CUDA capable graphics card; CUDA toolkit 4.2+; 6 GB global memory; 49 KB shared memory
Audience
Science/Research
This is an application that can also be fetched from https://sourceforge.net/projects/swiftseqaligner/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.