This is the Linux app named Taxoblast to run in Linux online whose latest release can be downloaded as Taxoblast1.21beta.jar. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named Taxoblast to run in Linux online with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start the OnWorks Linux online or Windows online emulator or MACOS online emulator from this website.
- 5. From the OnWorks Linux OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application, install it and run it.
SCREENSHOTS
Ad
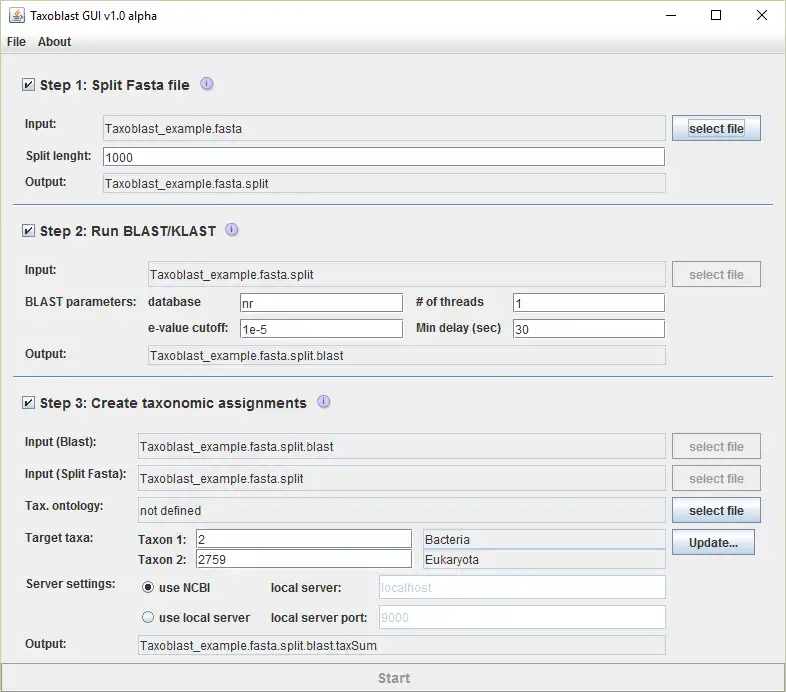
Taxoblast to run in Linux online
DESCRIPTION
Raw genomic sequences are frequently contaminated with sequences of other organism. Their identification is essential for the interpretation of genomic data. In this context it is essential to distinguish between horizontal gene transfers and contamination. The genomic context of sequences can help distinguish the two scenarios. Taxoblast splits genomic scaffolds into sub-sequences of defined length and for each of them determines the closest related taxon. It then summarizes this information for the entire scaffold, taking into account the taxonomic ontology. Scaffolds that exclusively match potential contaminants may be safely removed while sequences matching partially contaminants and partially the target organism may constitute horizontal transfers or assembly artifacts and need to be examined manually.Audience
Science/Research
User interface
Java Swing
Programming Language
Java
This is an application that can also be fetched from https://sourceforge.net/projects/taxoblast/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.