This is the Windows app named PARSEC - PAtteRn SEarch / Context to run in Windows online over Linux online whose latest release can be downloaded as PARSEC_Deployment_Package_v2.zip. It can be run online in the free hosting provider OnWorks for workstations.
Download and run online this app named PARSEC - PAtteRn SEarch / Context to run in Windows online over Linux online with OnWorks for free.
Follow these instructions in order to run this app:
- 1. Downloaded this application in your PC.
- 2. Enter in our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 3. Upload this application in such filemanager.
- 4. Start any OS OnWorks online emulator from this website, but better Windows online emulator.
- 5. From the OnWorks Windows OS you have just started, goto our file manager https://www.onworks.net/myfiles.php?username=XXXXX with the username that you want.
- 6. Download the application and install it.
- 7. Download Wine from your Linux distributions software repositories. Once installed, you can then double-click the app to run them with Wine. You can also try PlayOnLinux, a fancy interface over Wine that will help you install popular Windows programs and games.
Wine is a way to run Windows software on Linux, but with no Windows required. Wine is an open-source Windows compatibility layer that can run Windows programs directly on any Linux desktop. Essentially, Wine is trying to re-implement enough of Windows from scratch so that it can run all those Windows applications without actually needing Windows.
SCREENSHOTS
Ad
PARSEC - PAtteRn SEarch / Context to run in Windows online over Linux online
DESCRIPTION
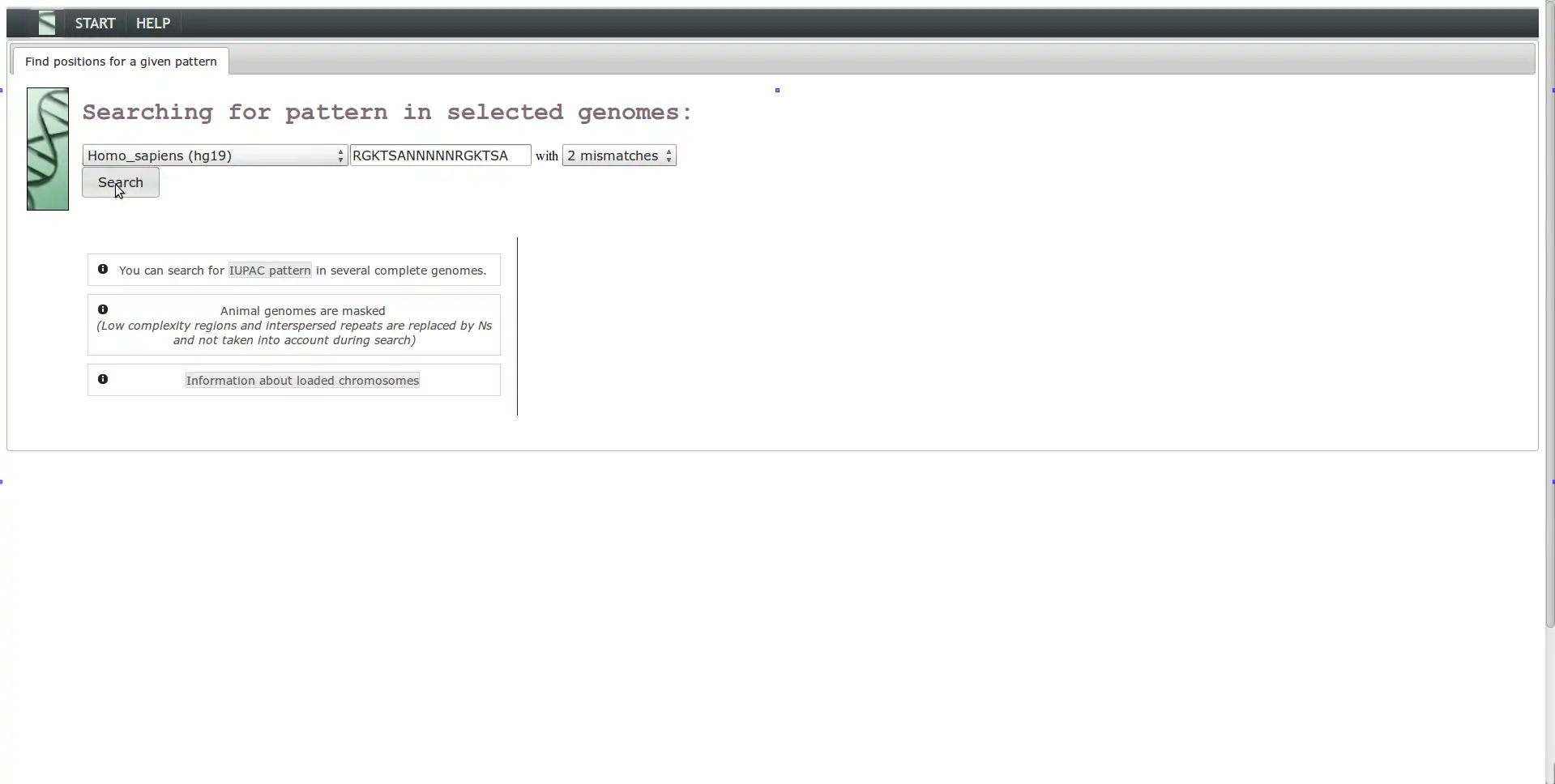
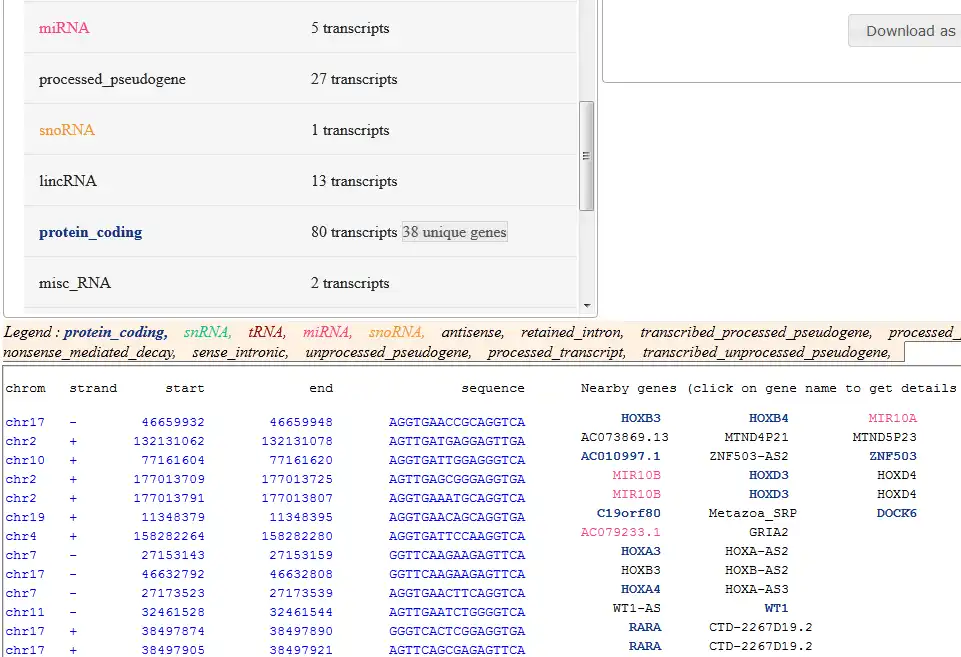
The characterization of genomic sites is a major challenge in the understanding and exploitation of next generation sequencing data. Most genomic sites are represented by short, degenerated motifs with a scattered distribution and sometimes with biological function (ex: regulation of gene expression, splicing patterns or epigenetics signals). These motifs are associated with a huge amount of noise and thus, the development of a computational platform for accurate detection of genomic sites requires the integration of various large-scale biological data in order to filter out false positives.PARSEC represents an intuitive, modular (easily extensible) and all-in-one solution for the efficient integration of lots of diverse genomic information in order to perform nonlinear localization and characterization of biological sites in a user-friendly environment.
See the wiki for hardware requirements and supported browsers.
Audience
Science/Research
User interface
Web-based
Programming Language
C++, Java
Database Environment
JDBC, MySQL, PostgreSQL (pgsql)
This is an application that can also be fetched from https://sourceforge.net/projects/genomicparsec/. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.